|
By Ian MacArthur
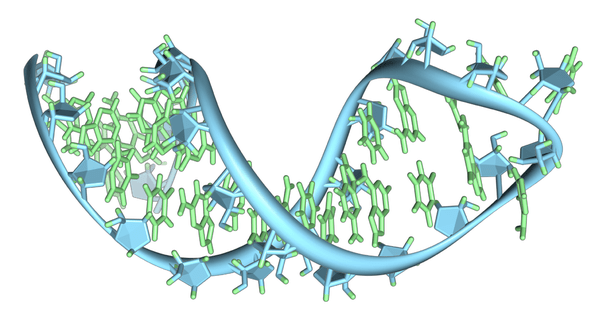
In biological sciences, researchers strive to understand life through observation. Attempts to artificially create life, however, can help further our mission to understand it. A team of scientists at Massachusetts General Hospital have overcome a major barrier in the replication of genetic material inside simple, artificial cell membranes—a feat that provides researchers with foundational information about how to create a primitive synthetic cell. Inspired by the challenge of simulating how the earliest cells functioned before the biological advent of enzymes, the research team led by Dr. Jack Szostak has been studying synthetic “protocells” comprised of simple fatty acid membranes containing RNA, the molecule believed to be the first nucleic acid used by cells to store genetic information. However, a major problem has long stood in the way of successfully creating a protocell capable of RNA copying: the magnesium ions required for RNA replication to proceed also disintegrate the fatty acid membrane that houses the genetic material. Without specialized enzymes of sufficient complexity to chemically protect cell membranes while catalyzing RNA replication, the earliest cells would have had to bypass this problem another way. The Szostak team believes they have identified a way that early cells might have done this. The researchers tested the ability of several chelating agents, or molecules that bind metal ions, to protect fatty acid membranes while preserving the chemical function of magnesium ions in RNA copying. To do this, they placed RNA molecules consisting of short primer sequences attached to longer template RNA strands inside protocell membranes. The single stranded portion of the template RNA strand contained a repeated sequence of cytosine nucleotides. Because the guanine nucleotide pairs with cytosine in double stranded nucleic acids, a successful chelating agent would allow the replication of the template strand to proceed if the cells were placed in a solution containing guanine. Citrate, an anionic derivative of citric acid, was found to be the most successful chelator in facilitating RNA replication. Dr. Szostak speculated as to how the molecule may have done this: “While other molecules can protect membranes against the magnesium ion, they don’t let RNA chemistry go on. We think that citrate is able both to protect membranes and to allow RNA copying to proceed by covering only one face of the magnesium ion, protecting the membrane while allowing RNA chemistry to work.” He later noted, however, that citrate would not have been abundant enough in the primeval Earth for early cells to have utilized it for this purpose. Although citrate may not have been the solution to non-enzymatic nucleic acid replication in the early Earth, it certainly provides a solution to the problem in the laboratory. The work of Dr. Szostak and his team are part of a larger effort by scientists to replicate biological function in the lab without the use of complex enzymes and other structures. The discovery and synthesis of molecules that mimic enzyme function is a major field with important implications for medicine and pharmacology. Since many biological compounds with important functions are too complex to synthesize on a large scale, it is the task of scientists to identify simpler molecules that can serve the same purpose. While Szostak’s research is not explicitly driven to find functional substitutes for complex molecules, it nevertheless demonstrates that tasks nominally performed by enzymes can be done by simpler compounds. This demonstration alone sustains hope that man may someday successfully synthesize a functional cell, which may further elucidate our cellular history.
0 Comments
By Ian MacArthur
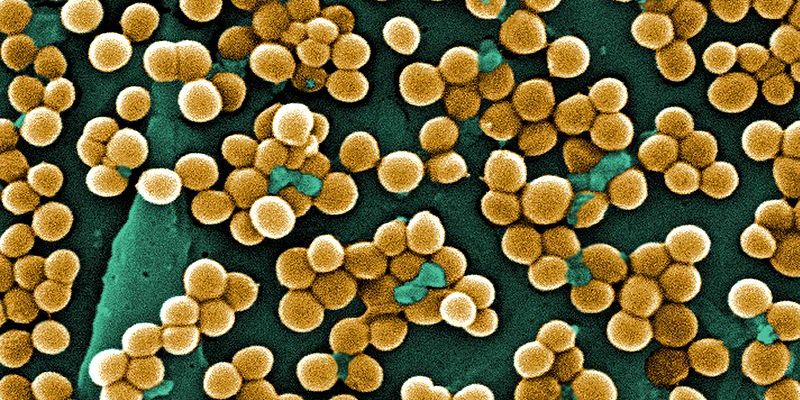
Antibiotic resistance in bacteria poses a major problem to modern public health. Infections that were once routinely treated with antibiotics have become extremely problematic to effectively kill, making even simple surgical operations dangerous from the perspective of contracting a resistant infection. Resistant strains of bacteria, such as methicillin-resistant Staphylococcus aureus, have sprung up in hospitals where they must be carefully contained lest the infection spread further. However, a new drug being developed by U.K. pharmaceutical company Helperby Therapeutics may open the door for methicillin and other antimicrobial agents to renew their therapeutic benefits. Researchers at the drug company have announced that they have identified a compound called HT61 that functions as an “antibiotic resistance breaker.” HT61, when paired with an antibiotic drug that a strain of bacteria have become resistant to, rejuvenates the effectiveness of the old antibiotic to kill colonies of the bacteria. Although the drug is not yet commercially available, a successful phase II clinical trial, in which scientists test whether the drug can be used in medical treatment, has paved the way toward cooperation between Helperby and an Indian drug company, Cadila Pharmaceuticals, to carry out a phase III trial. Investigators observed in the phase II trial that HT61 helped to dissolve the bacterial cell membrane, conferring new effectiveness to an antibiotic that a particular strain of staphylococcus had become resistant to. The identification of a resistance-breaking agent will no doubt inspire further research into compounds that may function similarly to restore the antimicrobial action of old antibiotics. Pending approval after the phase III trial, HT61 will likely play a large role in treating the existing resistant strains so as to mitigate the widespread problem of these strains appearing in hospitals. However, it is clear that drugs of this kind will not be a permanent solution to the problem of antibiotic resistance. If bacteria can become resistant to antimicrobial agents, there is no reason that, over time, they could not become resistant to resistance breakers. Indeed, it would take just a single gene mutation for a bacterium to achieve the means to counter the action of HT61. Once such a bacterium appears in a colony being treated with the drug, natural selection will select for bacteria with resistance to HT61 and the problem of resistance will persist. Anthony Coates, Chief Science Officer at Helperby, concisely summarized the current global problem of antibiotic resistance: “The emergence and spread of drug-resistant pathogens has accelerated whilst the pipeline for new anti-microbial drugs has all but run dry.” Implicit in this statement is the idea that intensified development of antibiotics may serve as a solution to drug resistance. While new drugs are certainly needed to treat the current problem in hospitals, it seems that using drugs to fight drug-resistant bacteria will lead to a pharmacological arms race between bacteria and humankind that man may not be posed to win. What will happen once bacteria become resistant to a new generation of antibiotics? Is the number of undiscovered antimicrobial compounds exhaustible? Or is there some other way to fight drug resistant infections besides treatment with other drugs? These are among the questions that should be asked as a way of determining how the world should move forward with the problem of antibiotic resistant bacteria. What is known now is that conventional methods will not be sufficient to finally solve the problem and that an immense amount of human imagination will be required to do so. Mankind will have to be diligent in formulating new ways of taking on this issue if our relative mastery of communicable diseases is to continue. By Alexandra DeCandia
Popcorn and Caramel breathed easily this Thanksgiving. The two lucky turkeys chosen for the annual Presidential pardon observed the holiday at the White House both feathered and free from the indignities of stuffing. However, as the winner and runner-up of a turkey equivalent to the Hunger Games (as Obama quipped), Popcorn and Caramel are fortunate to share such an unusual fate. Every year, Americans consume an estimated 46 million turkeys on the Thanksgiving holiday – almost one-fifth of the annual U.S. production of 254 million birds. Such astronomical, gravy-soaked casualties reveal more than an American zeal for turkey. It’s become a celebratory obsession – a staple during those times of family, warmth, and togetherness. But how much do Americans really know about the coveted drumsticks and over-sized breasts they consume each year? A short trip through the wormholes of the Internet reveals more than just basting techniques and optimal cooking times. The modern North American turkey (Meleagris gallopavo) descended from a long line of feathered bipeds that originated in the Early Miocene (c. 23 million years ago). Belonging to the order of Galliformes, they are most closely related to grouse, quail, pheasants, partridges, and chickens, but maintain the largest body size. Equipped with elaborate plumage (capable of shining red, purple, green, and gold in males), a varied repertoire of gobbles, and a distinctly nubby red head (complete with the ever attractive wattle, snood, and caruncles), wild turkeys aren’t exactly the subtlest of creatures. Therefore, it was not long before man, that great tinker of life, sought its domestication. Archaeological evidence suggests that turkeys were first domesticated by the Mesoamerican indigenes roughly 2,000 years ago. Likewise a staple for native peoples in the North, wild and newly domesticated turkeys provided ample eggs, meat, and decorative feathers for centuries. However, with European colonization of the Americas came the export and ultimate industrialization of the turkey industry. More and more, these intense selective pressures led domesticated turkeys to display phenotypes altered from their wild cousins. They doubled in size, developed white feathers, and grew such large breast muscles that they lost the ability of flight (and even that of natural copulation). Perhaps their greatest divergence, though, was one of population size. While the domesticated turkey population exploded to meet the ever-growing demand of a burgeoning nation, that of wild turkeys greatly diminished due to gross overhunting and rampant logging that coupled America’s economic expansion. By the early 1900’s, wild turkey populations reached an all-time low of 30,000 individuals. Restricted to isolated pockets of their ancestral range and almost entirely extirpated from its extremes, the species seemed en route to extinction. However, through a combination of strict hunting regulations, frequent individual relocations, and newly acquired funds from the 1937 Pittman-Robertson Act (which reallocated money from firearm taxes to wildlife conservation), wild turkey populations were able to rebound at an exponential rate. Today, more than 7 million wild turkeys join the 254 million domesticated turkeys currently inhabiting the North American continent. Their numbers are strong, their range expanding, and their meat so delicious when paired with good company and the perfect stuffing. Even Ben Franklin realized the significance of such a distinctly American bird. As he penned in a letter to his daughter (leading to the apocryphal myth of his desire for a Galliform national bird): “The turkey is…a respectable bird, and withal a true original Native of America…He is besides, though a little vain and silly, a Bird of Courage.” So be mindful and proud of your turkey next Thanksgiving. Remember the awkward majesty of its appearance, the struggle of its existence, and the tradition of its place at the center of American life and celebration. Then, once all due respect has been paid, gobble it up with the greatest care and ample amounts of gravy. |
Categories
All
Archives
April 2024
|