|
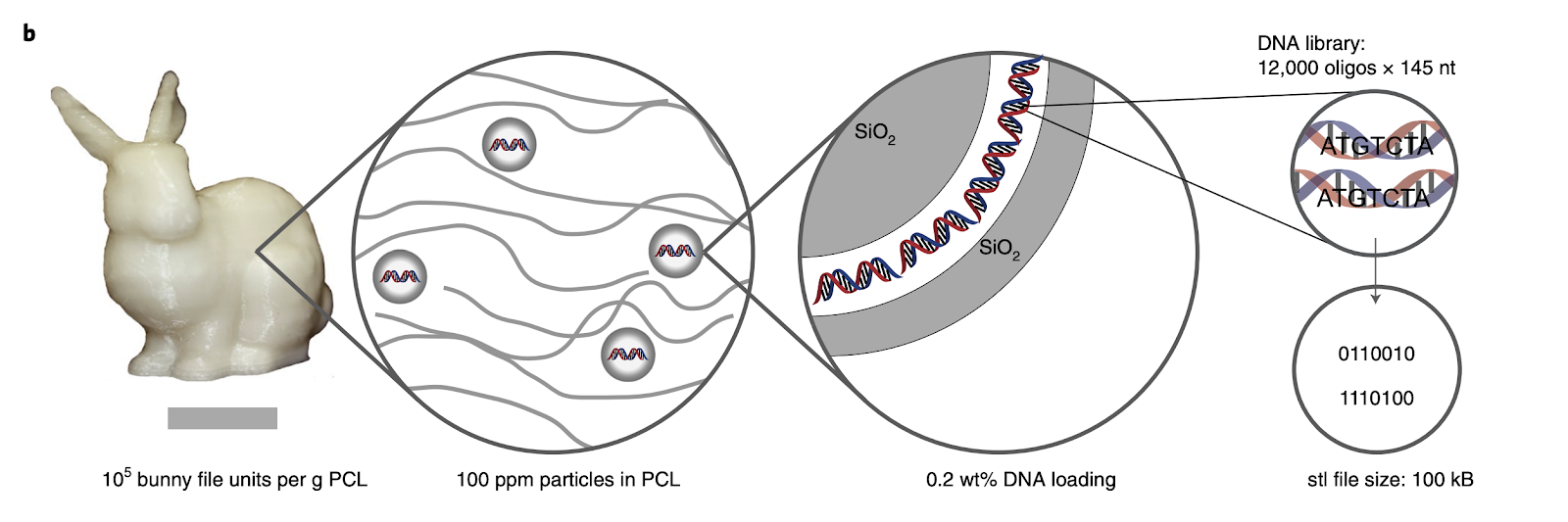
By Jenna Everard We all have digital data—texts, emails, essays, photos... the list goes on and on! And we all need to store it. To do so, we have flash drives, hard drives, servers, entire rooms lined with servers, and entire buildings filled with servers. The International Data Corporation has predicted that by 2025, the “global datasphere,” an entity that encompasses all the data created, captured, and replicated, will be around 175 Zettabytes, or 175 trillion gigabytes, in size. For perspective, if all that data was stored on DVDs, the stack of these DVDs would stretch around the Earth 222 times! With such rapid growth in data, it is likely that our infrastructure and capacity to store it will be tested. But what if the solution for doing so was just at our fingertips? Yes, literally at our fingertips, because the solution might just be our DNA. Characteristically, DNA is a fairly ideal storage device. While traditional floppy disks or flash drives are physically restricted to a rigid structure, DNA can exist as a liquid, allowing it to be inserted into practically any object. DNA is also very stable, so much so that scientists successfully sequenced genetic information from the fossil of a 700,000 year old horse. Comparably, common physical data-storage devices typically have lifespans of around a decade, and although the novel M-DISC claims to have a lifespan of 1,000 years, that is still miniscule when compared to DNA. Arguably the most significant advantage, though, is that DNA is capable of storing information at a very high density. In its microscopic structure of tightly wound double helices, the global datasphere would no longer stretch around the Earth 222 times. Instead, it could simply fit into the trunk of your car. Though this concept may appear to be an application of advanced modern biotechnology, it was actually first proposed over 50 years ago, before the first personal computer (the Kenbak-1) was even sold in 1971 and before the World Wide Web was released in 1991. American theoretical physicist Richard Feynman toyed with the idea of biological data storage in a well known talk of his in 1959, and Soviet physicist Mikhail Neiman specifically identified the potential for using DNA in 1964. Since these initial theorizations, many scientists, organizations, and corporations have tirelessly worked to bring these ideas to fruition. One of the early implementations of data storage in DNA was carried out in 1999 by a group of scientists at New York’s Mount Sinai School of Medicine. Inspired by the microdot technique used by German spies during World War II, where messages containing text and images were scaled to the size of a period and placed on objects ranging from letters to china dolls, these researchers aspired to create an even more discrete method of communication, this time using DNA. To do so, they leveraged a pre-existing structural feature of DNA: its organization into codons. A strand of DNA consists of four distinct nitrogen bases: Adenine (A), Thymine (T), Cytosine (C), and Guanine (G). In reading DNA, these bases are aggregated into groups of three, with each group representing one of 64 possible codons. In a biological system, each codon codes for an amino acid, and a chain of these amino acids ultimately forms a protein. Here, however, the researchers used each of the 64 codons to represent a letter, number, or symbol, and encoded their message “June 6 Invasion: Normandy”—alluding to the D-Day operation during World War II—by assembling the respective codons into a strand of DNA. They then, in a truly spy-like fashion, left traces of this DNA on a letter and sent it by mail. The recipients were then able to successfully recover their DNA and decode the original message. Since this implementation, the field of research surrounding digital data storage in DNA has continued to grow, and a major player in its expansion has been the Swiss Federal Institute of Technology in Zurich, more commonly referred to as ETH Zurich. In 2015, a team of researchers at ETH Zurich led by Robert Grass embedded data-containing DNA into silica nanobeads. Storing them in a heated environment of approximately 150 degrees Fahrenheit, they simulated the DNA breakdown that would happen over a few hundred years in a period of only a few weeks. From their results, they were able to formulate that DNA-encoded information would be able to persist for millions of years, shedding light on its stability and longevity. Four years later, in 2019, Robert Grass and his team of researchers, now also working with Yaniv Erlich (a former professor at Columbia University) unveiled their so-called DoT, short for “DNA of Things.” If you’re familiar with the IoT, or Internet of Things, then you’ll recognize this play on words. Just as the IoT describes how all of our devices are interconnected to exchange data and other information over the internet, the DoT describes a similar interconnectivity, except one in which data is exchanged through DNA. To translate the data into DNA, the researchers used a similarly nitrogen base focused approach, yet one that was fundamentally different in the manner it mapped the data. Essentially, the process went like this: the data to be stored was divided into smaller segments, after which a Luby Transformation was applied to each segment, encoding the data and producing blocks of 6 bits. Bits, short for binary digits, are the smallest units of measurement of computer data, containing only the binary values of 0 and 1. To begin converting these blocks of bit sequences into DNA sequences, the bits were grouped into pairs, of which there are only four possible pairings: 00, 01, 10, and 11. Coincidentally, there are also only four nucleotide bases in DNA, hence each pair of bits could be converted into a base, and, ultimately, the string of bits could be converted into a DNA sequence. Once this sequence was obtained, the DNA was then embedded into the silica nanobeads, which allowed it fused into practically any material. Their choice application of their new technology was to 3D print a Stanford Bunny, a common model for testing computer graphics. What made this Stanford Bunny unique though, was that embedded in its synthetic polyester makeup were DNA molecules that contained the exact instructions for re-printing the bunny. As outlined in their paper, the ETH Zurich researchers envision a wide array of future uses for their DoT, from storing a patient’s medical records within their medical implants to building machines that are capable of self-replication. Now, just three months ago, it was announced that ETH Zurich had joined the new DNA Data Storage Alliance, originally founded by companies including Microsoft and Illumina. Over the past two decades, the alliance’s members have worked on numerous independent projects, manipulating DNA to store larger and larger amounts of data. They have stored the movie The Wizard of Oz, the entirety of War and Peace, Amazon gift cards, a complete computer operating system, and even a computer virus (which, when sequenced, actually allowed the original researchers to take over the computer that became infected with the malware!). Though the current high price of DNA digital data storage limits its everyday practicality, there is much hope that this collaboration will lead to great strides in the efficiency, capabilities and applications of storing our data in DNA. So the next time you’re left grumbling after your flash drive conveniently decides to corrupt the night before your final presentation, just remember that you are the solution. With our own DNA we have the potential to revolutionize our data storage mechanisms and ensure we can support and securely store the continually growing global datasphere. Yet this begs the question then, with the exponential growth of the global datasphere, what happens when it surpasses even the data storage capacity of our DNA?
0 Comments
Leave a Reply. |
Categories
All
Archives
April 2024
|